Interpreting model results¶
After calling interlace.fit(), you receive a CrossedLMEResult object. This page
explains what each part of the output means and how to use it to draw conclusions from
your model.
Throughout this page we use a running example: exam scores modelled as a function of hours studied and prior GPA, with students and schools as crossed grouping factors.
import numpy as np
import pandas as pd
import interlace
rng = np.random.default_rng(42)
n = 300
df = pd.DataFrame({
"score": 60 + rng.normal(0, 8, n),
"hours_studied": rng.uniform(0, 10, n),
"prior_gpa": rng.uniform(2.0, 4.0, n),
"student_id": [f"s{i}" for i in rng.integers(1, 31, n)],
"school_id": [f"sch{i}" for i in rng.integers(1, 11, n)],
})
result = interlace.fit(
"score ~ hours_studied + prior_gpa",
data=df,
groups=["student_id", "school_id"],
)
/tmp/ipykernel_2509/4095886218.py:16: UserWarning: boundary (singular) fit: see help(interlace.isSingular)
Fixed effects¶
Fixed effects capture the population-level relationships between predictors and the outcome, controlling for the grouping structure.
The result exposes four attributes for fixed-effect inference:
Attribute |
Contents |
|---|---|
|
Coefficient estimates ( |
|
Standard errors |
|
Two-sided p-values (normal approximation) |
|
95% confidence intervals ( |
Reading the coefficients¶
fe_params is a pandas.Series indexed by term name. Each coefficient is the
expected change in the outcome per unit increase in the predictor, holding grouping
structure constant — i.e., the within-group effect.
result.fe_params
Intercept 60.454607
hours_studied 0.032509
prior_gpa -0.322930
dtype: float64
Standard errors and inference¶
Standard errors in fe_bse account for the clustering: they are derived from the
fixed-effect covariance matrix fe_cov = scale * (X'Ω⁻¹X)⁻¹, where X is the
fixed-effect design matrix (one row per observation, one column per predictor including
the intercept) and Ω is the marginal covariance matrix of the response that encodes the
random effect structure. This is why they are typically larger than OLS standard errors
on the same data — the effective sample size is reduced by the grouping.
A convenient summary table:
pd.DataFrame({
"estimate": result.fe_params,
"se": result.fe_bse,
"p": result.fe_pvalues,
"ci_lo": result.fe_conf_int.iloc[:, 0],
"ci_hi": result.fe_conf_int.iloc[:, 1],
})
| estimate | se | p | ci_lo | ci_hi | |
|---|---|---|---|---|---|
| Intercept | 60.454607 | 2.446897 | 0.000000 | 55.658688 | 65.250526 |
| hours_studied | 0.032509 | 0.149269 | 0.827601 | -0.260059 | 0.325077 |
| prior_gpa | -0.322930 | 0.714485 | 0.651300 | -1.723321 | 1.077460 |
P-values use a normal approximation (z-test), which is appropriate for large samples. For small samples with few groups, consider using likelihood-ratio tests or parametric bootstrap instead.
Comparing models with different fixed structures¶
To test whether a predictor improves fit, refit with method="ML" (not REML) and
compare log-likelihoods. Use REML (the default) for your final reported estimates;
use ML only for likelihood-ratio tests comparing fixed structures. See
Concepts: REML for the reason.
from scipy.stats import chi2
m1 = interlace.fit("score ~ hours_studied", data=df, groups=["student_id", "school_id"], method="ML")
m2 = interlace.fit("score ~ hours_studied + prior_gpa", data=df, groups=["student_id", "school_id"], method="ML")
lr_stat = 2 * (m2.llf - m1.llf)
df_diff = 1 # one extra parameter in m2
p_value = chi2.sf(lr_stat, df=df_diff)
print(f"LR statistic: {lr_stat:.3f} (df={df_diff})")
print(f"p-value: {p_value:.4f}")
LR statistic: 0.210 (df=1)
p-value: 0.6470
/tmp/ipykernel_2509/2957097658.py:3: UserWarning: boundary (singular) fit: see help(interlace.isSingular)
/tmp/ipykernel_2509/2957097658.py:4: UserWarning: boundary (singular) fit: see help(interlace.isSingular)
Variance components¶
Variance components quantify how much variability in the outcome is attributable to each grouping factor and to residual within-group noise.
Reading variance components¶
print("Between-group variances:")
for g, v in result.variance_components.items():
print(f" {g}: σ² = {v:.3f} (σ ≈ {v**0.5:.2f} score points)")
print(f"\nResidual variance (scale): σ² = {result.scale:.3f} (σ ≈ {result.scale**0.5:.2f} score points)")
Between-group variances:
student_id: σ² = 4.513 (σ ≈ 2.12 score points)
school_id: σ² = 0.000 (σ ≈ 0.00 score points)
Residual variance (scale): σ² = 51.391 (σ ≈ 7.17 score points)
Intraclass correlation (ICC)¶
The ICC tells you what fraction of total variance each grouping factor explains. An ICC below ~0.05 suggests the grouping factor contributes little and may not need a random intercept.
total = sum(result.variance_components.values()) + result.scale
icc = {g: v / total for g, v in result.variance_components.items()}
for g, v in icc.items():
print(f"ICC({g}): {v:.3f} → {v*100:.1f}% of variance explained by {g}")
ICC(student_id): 0.081 → 8.1% of variance explained by student_id
ICC(school_id): 0.000 → 0.0% of variance explained by school_id
What if a variance component is near zero?¶
A variance component close to zero (sometimes reported as exactly 0 at the boundary of the parameter space) means the corresponding grouping factor explains almost no variance in the outcome after conditioning on the fixed effects. This is not an error — it is a valid result indicating that the groups in that factor are essentially homogeneous. You may choose to drop that grouping factor and refit, or retain it for theoretical reasons.
Model fit¶
print(f"AIC: {result.aic:.2f}")
print(f"BIC: {result.bic:.2f}")
print(f"log-lik: {result.llf:.2f}")
print(f"converged: {result.converged}")
AIC: 2061.00
BIC: 2083.22
log-lik: -1024.50
converged: True
AIC and BIC¶
AIC and BIC are used to compare models with the same fixed structure fitted with REML, or models with different fixed structures fitted with ML. Both penalise model complexity; BIC penalises more heavily for large samples.
AIC is defined as −2 log L + 2k (Akaike, 1974); BIC as −2 log L + k log n (Schwarz, 1978), where k is the number of estimated parameters and n the number of observations.
A difference of ≥ 4 AIC points between models is often treated as meaningful; a difference of 1–3 is weak evidence. These are heuristics — use them alongside subject-matter reasoning, not as automatic decision rules.
References
Akaike, H. (1974). A new look at the statistical model identification. IEEE Transactions on Automatic Control, 19(6), 716–723. doi:10.1109/TAC.1974.1100705
Schwarz, G. (1978). Estimating the dimension of a model. The Annals of Statistics, 6(2), 461–464. doi:10.1214/aos/1176344136
Checking convergence¶
Always verify convergence before reporting results. Non-convergence typically occurs with very small datasets, degenerate variance (one group per level), or near-zero variance components.
if not result.converged:
print("Warning: optimiser did not converge — estimates may be unreliable")
else:
print("Converged successfully.")
Converged successfully.
Random effects (BLUPs)¶
BLUPs (Best Linear Unbiased Predictors) are the estimated deviations of each group
from the population mean. They are accessible per grouping factor as a dict of
pd.Series (intercept-only models) or pd.DataFrame (random slopes models).
Reading BLUPs¶
student_blups = result.random_effects["student_id"]
school_blups = result.random_effects["school_id"]
student_blups.sort_values()
s17 -2.813086
s12 -2.566059
s18 -1.792370
s9 -1.789333
s3 -1.499837
s22 -1.007045
s11 -0.940747
s16 -0.891217
s21 -0.824995
s6 -0.657869
s19 -0.594837
s28 -0.518625
s2 -0.512487
s7 -0.369438
s14 -0.158327
s30 0.032735
s29 0.120162
s26 0.324681
s5 0.325169
s20 0.363378
s8 0.379302
s23 0.605424
s15 0.829710
s1 1.317096
s13 1.367523
s27 1.627519
s24 1.823487
s4 2.066743
s25 2.825764
s10 2.927580
Name: student_id, dtype: float64
Each value is the estimated deviation of that group from the population mean, in the same units as the outcome (exam score points). A positive BLUP means the group scores above average after accounting for the fixed predictors and other grouping factors.
For random slopes models, random_effects[group] is a pd.DataFrame with one column
per term (intercept + slope).
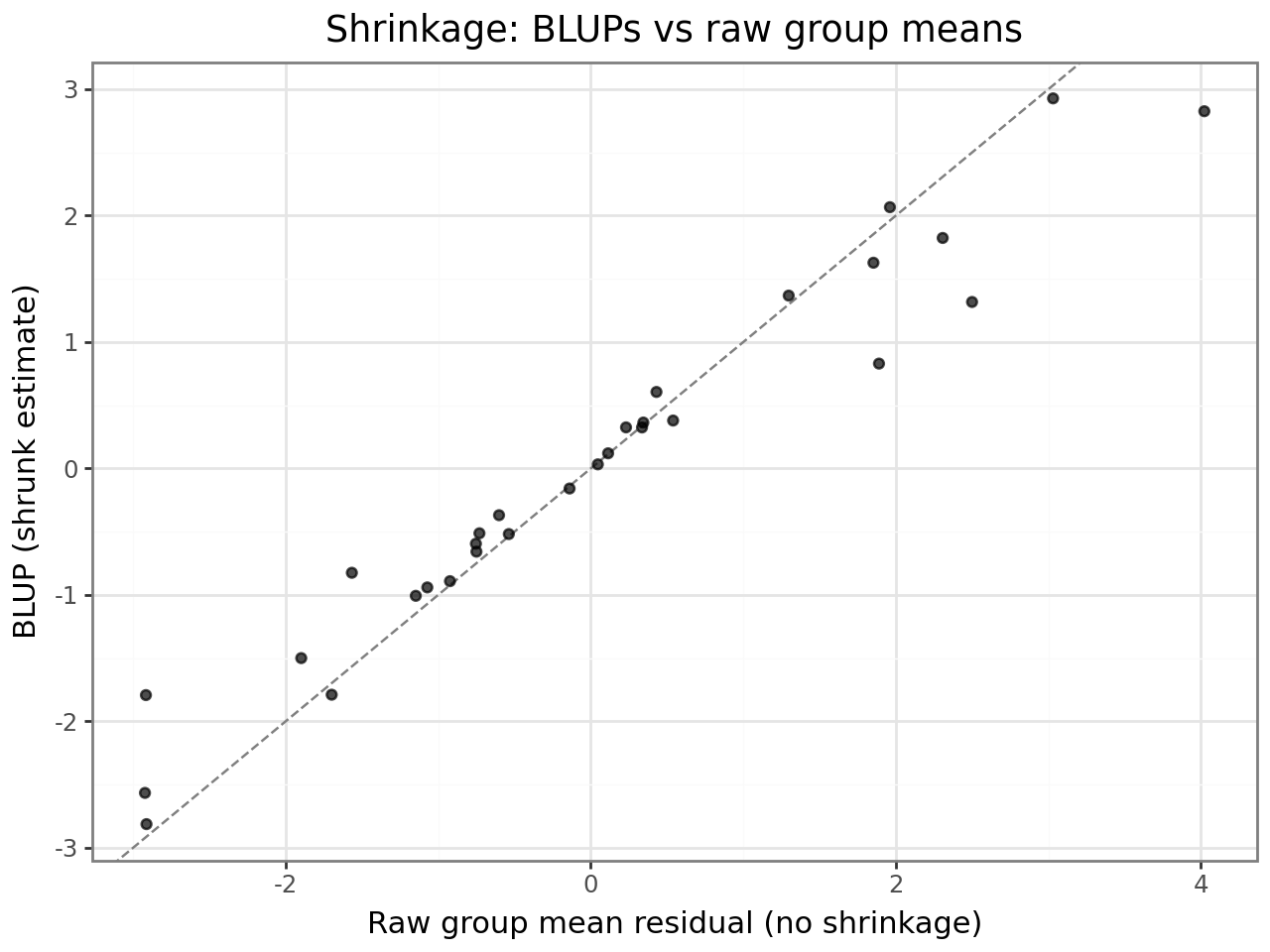
Shrinkage¶
BLUPs are shrunk toward zero relative to the group’s raw deviation. Groups with fewer observations or groups whose variance component is small are shrunk more. This is intentional: a student with only one observation should not be assigned an extreme intercept — the BLUP borrows strength from the population distribution.
The plot below makes shrinkage visible: each point is one student. Points above the diagonal are shrunk toward zero; the further from the line, the stronger the shrinkage.
from plotnine import ggplot, aes, geom_point, geom_abline, labs, theme_bw
raw = df.assign(resid=result.resid).groupby("student_id")["resid"].mean()
shrinkage_df = pd.DataFrame({
"raw_mean_resid": raw,
"blup": student_blups.reindex(raw.index),
}).reset_index()
(
ggplot(shrinkage_df, aes(x="raw_mean_resid", y="blup"))
+ geom_abline(slope=1, intercept=0, linetype="dashed", color="grey")
+ geom_point(alpha=0.7)
+ labs(
x="Raw group mean residual (no shrinkage)",
y="BLUP (shrunk estimate)",
title="Shrinkage: BLUPs vs raw group means",
)
+ theme_bw()
)
BLUPs vs fixed group dummies¶
BLUPs are not the same as including a fixed dummy variable for each group. Dummies use only the data from that group; BLUPs partially pool across groups. For groups with very many observations, BLUPs converge to the dummy-variable estimate. For small or sparse groups, BLUPs are much more stable.
Predictions¶
result.predict() returns fitted values, optionally including random effects.
Conditional ( |
Marginal ( |
|
|---|---|---|
Includes BLUPs |
Yes |
No |
Use case |
In-sample fit, known groups |
New groups, population-level |
Unseen group levels |
BLUP = 0 (shrinks to mean) |
Identical to marginal |
Conditional predictions are appropriate when you know which group an observation
belongs to and want the best estimate for that specific group. They are used to
compute residuals (result.resid = observed − conditional prediction).
Marginal predictions give the expected outcome for a hypothetical average group — useful for reporting population-level effects or when the new data includes groups not seen during training.
# Conditional predictions (uses BLUPs, default)
y_conditional = result.predict()
# Marginal predictions (fixed effects only)
y_marginal = result.predict(include_re=False)
print(f"Conditional predictions — mean: {y_conditional.mean():.2f}, std: {y_conditional.std():.2f}")
print(f"Marginal predictions — mean: {y_marginal.mean():.2f}, std: {y_marginal.std():.2f}")
Conditional predictions — mean: 59.67, std: 1.43
Marginal predictions — mean: 59.67, std: 1.43
Predicting on new data¶
Unseen group levels automatically receive a BLUP of zero — the population mean for that factor. This is the correct Bayesian-optimal behaviour under the model’s assumptions.
df_new = pd.DataFrame({
"hours_studied": [5.0, 8.0, 5.0],
"prior_gpa": [3.2, 3.8, 3.2],
"student_id": ["s1", "s2", "s_new"], # s_new is unseen → BLUP = 0
"school_id": ["sch1", "sch1", "sch1"],
})
preds = result.predict(newdata=df_new)
df_new.assign(predicted_score=preds.round(2))
| hours_studied | prior_gpa | student_id | school_id | predicted_score | |
|---|---|---|---|---|---|
| 0 | 5.0 | 3.2 | s1 | sch1 | 60.90 |
| 1 | 8.0 | 3.8 | s2 | sch1 | 58.98 |
| 2 | 5.0 | 3.2 | s_new | sch1 | 59.58 |
Bootstrap standard error¶
For scalar statistics of the outcome (currently: median), bootstrap_se() computes
a cluster-bootstrap standard error that respects the grouping structure.
Ordinary bootstrap resamples individual observations, which underestimates the SE when group variance is substantial. Cluster bootstrap resamples group levels with replacement, then includes all observations from the sampled groups — preserving within-group correlation.
Use resample_level="observation" only if you have verified that the ICC is negligible.
se_cluster = result.bootstrap_se(statistic="median", n_bootstrap=500, resample_level="group", seed=42)
se_observation = result.bootstrap_se(statistic="median", n_bootstrap=500, resample_level="observation", seed=42)
print(f"Median: {np.median(df['score']):.2f}")
print(f"Cluster-bootstrap SE: {se_cluster:.4f}")
print(f"Observation-bootstrap SE: {se_observation:.4f} (underestimates when ICC > 0)")
Median: 59.30
Cluster-bootstrap SE: 0.8252
Observation-bootstrap SE: 0.5958 (underestimates when ICC > 0)
Quick reference¶
Attribute |
Type |
What it tells you |
|---|---|---|
|
|
Fixed-effect coefficient estimates |
|
|
Standard errors of fixed effects |
|
|
Two-sided p-values (z-test) |
|
|
95% confidence intervals |
|
|
Fixed-effect covariance matrix |
|
|
Between-group variance per factor |
|
|
Within-group (residual) variance σ² |
|
|
BLUPs per grouping factor |
|
|
Conditional fitted values |
|
|
Conditional residuals |
|
|
Akaike Information Criterion |
|
|
Bayesian Information Criterion |
|
|
Log-likelihood at optimum |
|
|
Whether the optimiser converged |
|
|
Number of observations |
|
|
Number of levels per grouping factor |
Where to go next¶
Diagnostics guide — residuals, leverage, Cook’s D, influence plots
API: CrossedLMEResult — full attribute and method reference
API: Prediction —
predict()parameter detailsConcepts — background on REML, BLUPs, and variance components