Case Study: Cox Frailty Model for Kidney Stone Recurrence¶
This notebook walks through a realistic multi-center clinical trial analysis using interlace.coxme() — a Cox proportional hazards model with Gaussian frailty.
Why Cox frailty over standard Cox PH?¶
Standard Cox PH assumes all patients share the same baseline hazard, ignoring clustering. When patients are nested in centers (hospitals, clinics), unobserved center-level differences inflate the apparent treatment effect and produce overconfident standard errors. A frailty model adds a random intercept per center on the log-hazard scale, capturing this heterogeneity explicitly.
Feature |
Standard Cox |
Cox Frailty ( |
|---|---|---|
Handles clustering |
No |
Yes |
Center-level BLUPs |
No |
Yes |
Correct SE under clustering |
No |
Yes |
Variance components |
No |
Yes |
Dataset: Kidney Stone Recurrence Trial¶
300 patients across 20 clinical centers (15 per center)
Outcome: time-to-recurrence in months, right-censored
Predictors:
treatment(0=standard, 1=new),age_c(age mean-centred at 58, divided by 10)Random effect:
center(frailty, Gaussian on log-hazard scale)
import numpy as np
import pandas as pd
import interlace
rng = np.random.default_rng(99)
TRUE_LAMBDA0 = 0.04
TRUE_BETA_TRT = np.log(0.65) # -0.431
TRUE_BETA_AGE = np.log(1.25) # 0.223 (per 10 years)
TRUE_SD_CENTER = 0.5
n_centers = 20
n_per_center = 15
u_center = rng.normal(0, TRUE_SD_CENTER, n_centers)
rows = []
for j in range(n_centers):
for i in range(n_per_center):
trt = int(rng.binomial(1, 0.5))
age_raw = rng.normal(58, 12)
age_c = (age_raw - 58) / 10
h = TRUE_LAMBDA0 * np.exp(TRUE_BETA_TRT * trt + TRUE_BETA_AGE * age_c + u_center[j])
surv_time = rng.exponential(1 / h)
cens_time = rng.uniform(12, 36)
obs_time = min(surv_time, cens_time)
event = int(surv_time <= cens_time)
rows.append({
'patient': f'P{j*n_per_center+i+1:03d}',
'center': f'C{j+1:02d}',
'treatment': trt,
'age': round(age_raw, 1),
'age_c': round(age_c, 3),
'time': round(obs_time, 2),
'event': event,
})
df = pd.DataFrame(rows)
df['trt_label'] = df['treatment'].map({0: 'Standard', 1: 'New treatment'})
print(f"Observations : {len(df)}")
print(f"Centers : {df['center'].nunique()}")
print(f"Event rate : {df['event'].mean():.1%}")
print()
print("Event rate by treatment:")
print(df.groupby('trt_label')['event'].agg(['sum', 'mean']).rename(columns={'sum': 'events', 'mean': 'rate'}))
Observations : 300
Centers : 20
Event rate : 57.0%
Event rate by treatment:
events rate
trt_label
New treatment 72 0.464516
Standard 99 0.682759
df[['patient', 'center', 'treatment', 'age', 'time', 'event']].head(8)
| patient | center | treatment | age | time | event | |
|---|---|---|---|---|---|---|
| 0 | P001 | C01 | 0 | 53.8 | 33.36 | 0 |
| 1 | P002 | C01 | 0 | 39.8 | 14.98 | 1 |
| 2 | P003 | C01 | 1 | 57.0 | 8.43 | 1 |
| 3 | P004 | C01 | 0 | 50.3 | 1.64 | 1 |
| 4 | P005 | C01 | 1 | 68.1 | 13.38 | 1 |
| 5 | P006 | C01 | 0 | 75.7 | 25.74 | 0 |
| 6 | P007 | C01 | 1 | 78.4 | 30.41 | 1 |
| 7 | P008 | C01 | 1 | 54.1 | 2.87 | 1 |
Exploratory Data Analysis¶
Before fitting the model, we examine:
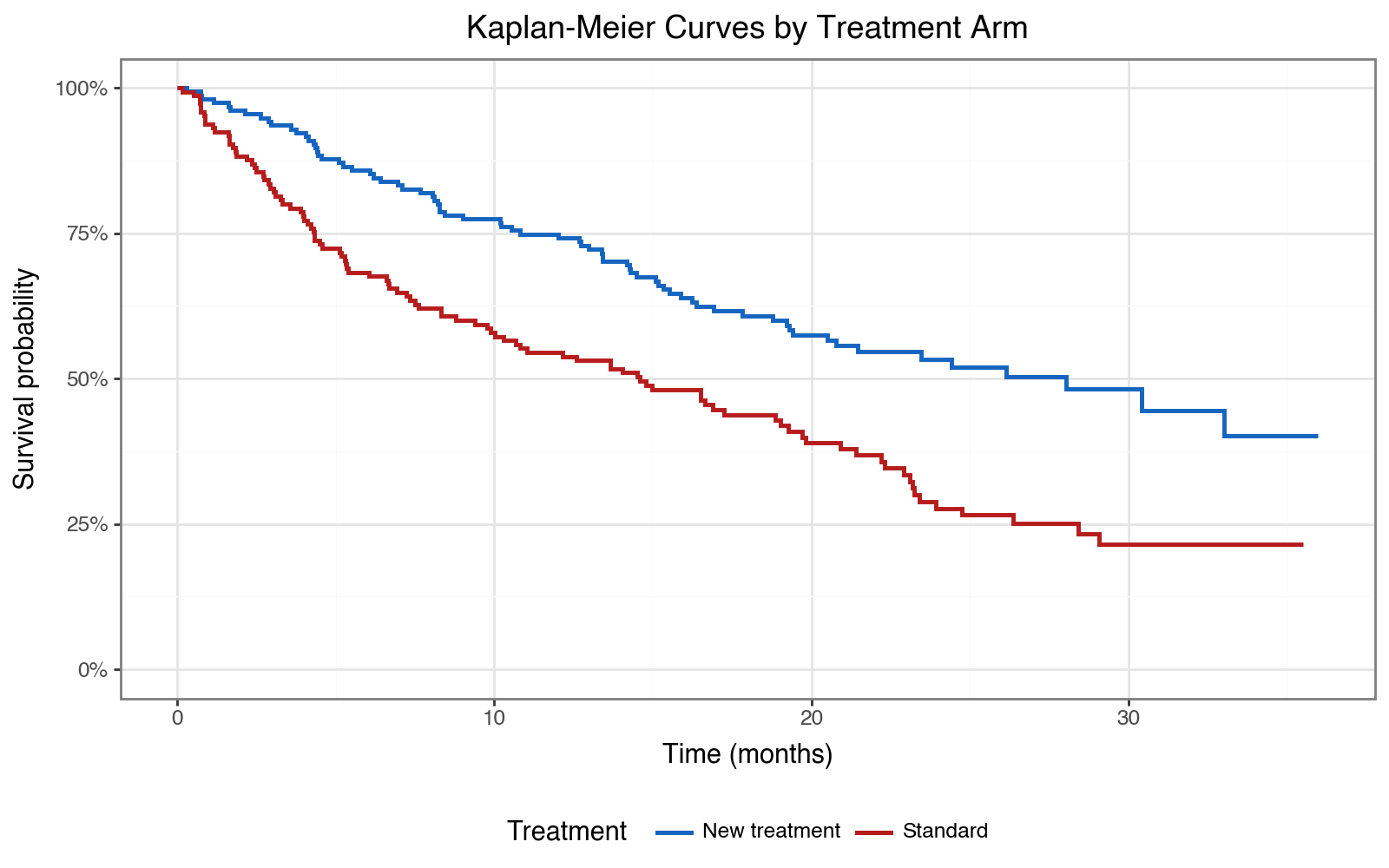
Kaplan-Meier curves by treatment arm — do the arms separate?
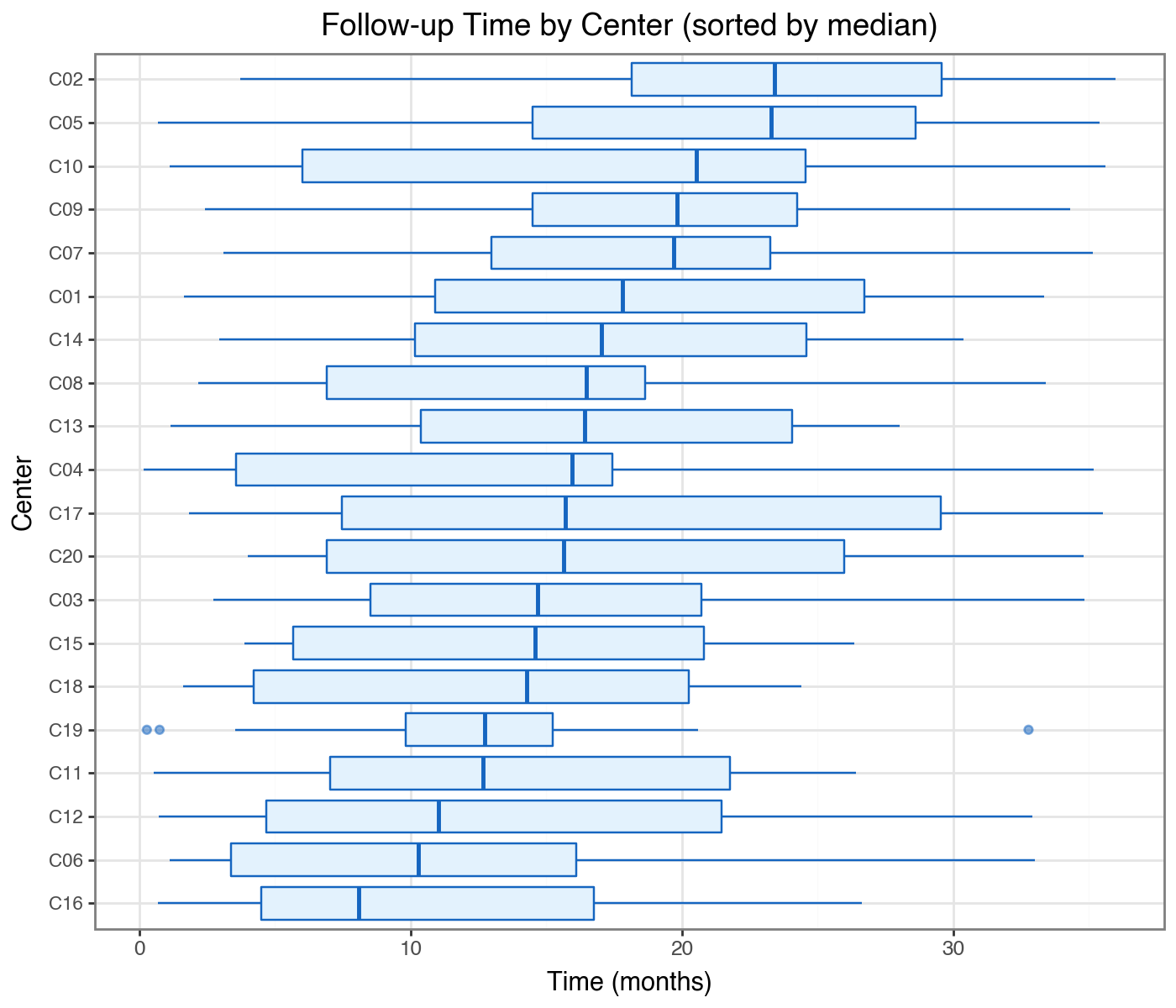
Follow-up time by center — is there between-center heterogeneity in censoring patterns?
from plotnine import (
ggplot, aes, geom_step, geom_point, geom_errorbarh, geom_errorbar,
geom_vline, geom_hline, geom_boxplot, coord_flip,
scale_color_manual, scale_x_continuous, scale_y_continuous,
labs, theme_bw, theme, element_text, element_blank, facet_wrap
)
def km_df(data, time_col, event_col, group_col):
rows = []
for g in sorted(data[group_col].unique()):
sub = data[data[group_col] == g].sort_values(time_col)
t, e = sub[time_col].values, sub[event_col].values
n = len(t)
surv = 1.0
rows.append({group_col: g, 'time': 0.0, 'survival': 1.0})
for i in range(n):
if e[i]:
surv *= (1 - 1 / (n - i))
rows.append({group_col: g, 'time': float(t[i]), 'survival': surv})
return pd.DataFrame(rows)
km = km_df(df, 'time', 'event', 'trt_label')
(
ggplot(km, aes(x='time', y='survival', color='trt_label'))
+ geom_step(size=1)
+ scale_color_manual(values={'Standard': '#B71C1C', 'New treatment': '#1565C0'})
+ scale_y_continuous(limits=(0, 1), labels=lambda l: [f'{v:.0%}' for v in l])
+ labs(
title='Kaplan-Meier Curves by Treatment Arm',
x='Time (months)',
y='Survival probability',
color='Treatment',
)
+ theme_bw()
+ theme(legend_position='bottom', figure_size=(8, 5))
)
center_order = (
df.groupby('center')['time']
.median()
.sort_values()
.index.tolist()
)
df['center_f'] = pd.Categorical(df['center'], categories=center_order, ordered=True)
(
ggplot(df, aes(x='center_f', y='time'))
+ geom_boxplot(fill='#E3F2FD', color='#1565C0', outlier_alpha=0.5)
+ coord_flip()
+ labs(
title='Follow-up Time by Center (sorted by median)',
x='Center',
y='Time (months)',
)
+ theme_bw()
+ theme(figure_size=(7, 6), axis_text_y=element_text(size=8))
)
Model Specification¶
We fit:
$$ h_{ij}(t) = h_0(t) \exp\bigl(\beta_{\text{trt}} \cdot x_{\text{trt}} + \beta_{\text{age}} \cdot x_{\text{age}} + u_j\bigr) $$
where $u_j \sim \mathcal{N}(0, \sigma^2_{\text{center}})$ is the frailty for center $j$.
The interlace.coxme() interface mirrors statsmodels.MixedLM.from_formula(): fixed effects go in formula, the grouping variable in groups.
result = interlace.coxme(
formula="Surv(time, event) ~ treatment + age_c",
data=df,
groups="center",
)
print(f"Converged : {result.converged}")
print(f"N obs : {result.nobs}")
print(f"Events : {result.n_events}")
print(f"AIC : {result.aic:.2f}")
print(f"BIC : {result.bic:.2f}")
Converged : True
N obs : 300
Events : 171
AIC : 1756.67
BIC : 1767.78
fe = result.fe_params
ci = result.fe_conf_int
bse = result.fe_bse
pval = result.fe_pvalues
hr_table = pd.DataFrame({
'log_HR': fe,
'HR': np.exp(fe),
'CI_lo': np.exp(ci['lower']),
'CI_hi': np.exp(ci['upper']),
'SE': bse,
'p_value': pval,
})
print("Hazard Ratio Table")
print(hr_table.round(4))
print()
print(f"Center frailty SD : {np.sqrt(result.variance_components['center']):.4f} (true={TRUE_SD_CENTER})")
print(f"True log HR trt : {TRUE_BETA_TRT:.4f} estimated: {fe.get('treatment', float('nan')):.4f}")
print(f"True log HR age : {TRUE_BETA_AGE:.4f} estimated: {fe.get('age_c', float('nan')):.4f}")
Hazard Ratio Table
log_HR HR CI_lo CI_hi SE p_value
treatment -0.6097 0.5435 0.3985 0.7413 0.1583 0.0001
age_c 0.0983 1.1033 0.9730 1.2511 0.0641 0.1252
Center frailty SD : 0.3360 (true=0.5)
True log HR trt : -0.4308 estimated: -0.6097
True log HR age : 0.2231 estimated: 0.0983
forest_df = hr_table.reset_index().rename(columns={'index': 'Parameter'})
(
ggplot(forest_df, aes(x='HR', y='Parameter'))
+ geom_vline(xintercept=1, linetype='dashed', color='grey')
+ geom_errorbarh(aes(xmin='CI_lo', xmax='CI_hi'), height=0.2, color='#1565C0')
+ geom_point(size=4, color='#B71C1C')
+ scale_x_continuous(limits=(0, 2))
+ labs(
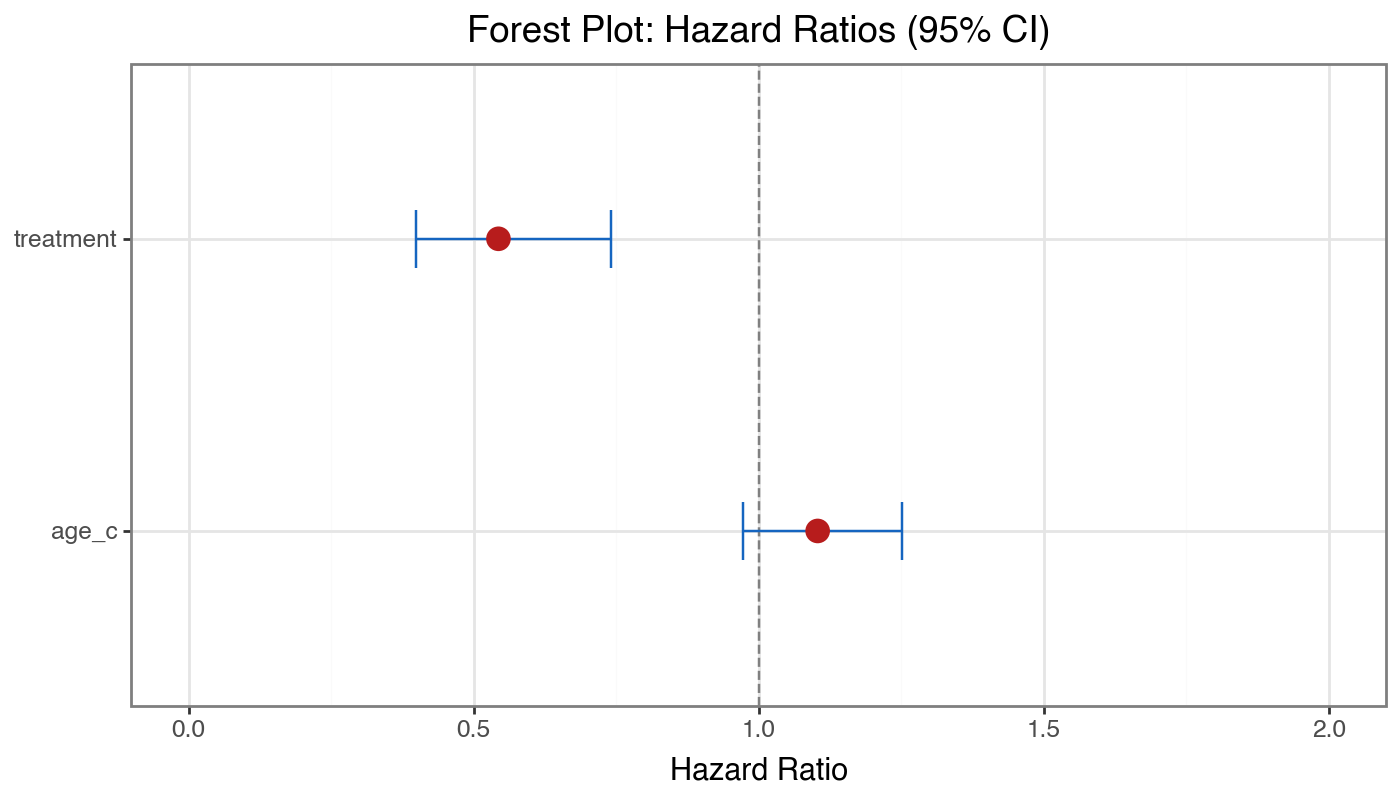
title='Forest Plot: Hazard Ratios (95% CI)',
x='Hazard Ratio',
y='',
)
+ theme_bw()
+ theme(figure_size=(7, 4))
)
print(f"Concordance (C-statistic): {result.concordance:.4f}")
print()
print("A C-statistic > 0.5 indicates better-than-chance discrimination.")
print("The frailty model accounts for center clustering; the population-averaged")
print("C-statistic reflects discrimination after integrating out random effects.")
Concordance (C-statistic): 0.6486
A C-statistic > 0.5 indicates better-than-chance discrimination.
The frailty model accounts for center clustering; the population-averaged
C-statistic reflects discrimination after integrating out random effects.
blups = result.random_effects['center'].sort_values()
blup_se = np.sqrt(result.variance_components['center'])
cat_df = pd.DataFrame({
'center': blups.index,
'blup': blups.values,
'lo': blups.values - 1.96 * blup_se,
'hi': blups.values + 1.96 * blup_se,
})
cat_df['center'] = pd.Categorical(cat_df['center'], categories=cat_df['center'].tolist(), ordered=True)
(
ggplot(cat_df, aes(x='center', y='blup'))
+ geom_hline(yintercept=0, linetype='dashed', color='grey')
+ geom_errorbar(aes(ymin='lo', ymax='hi'), width=0.3, color='#1565C0')
+ geom_point(size=3, color='#B71C1C')
+ coord_flip()
+ labs(
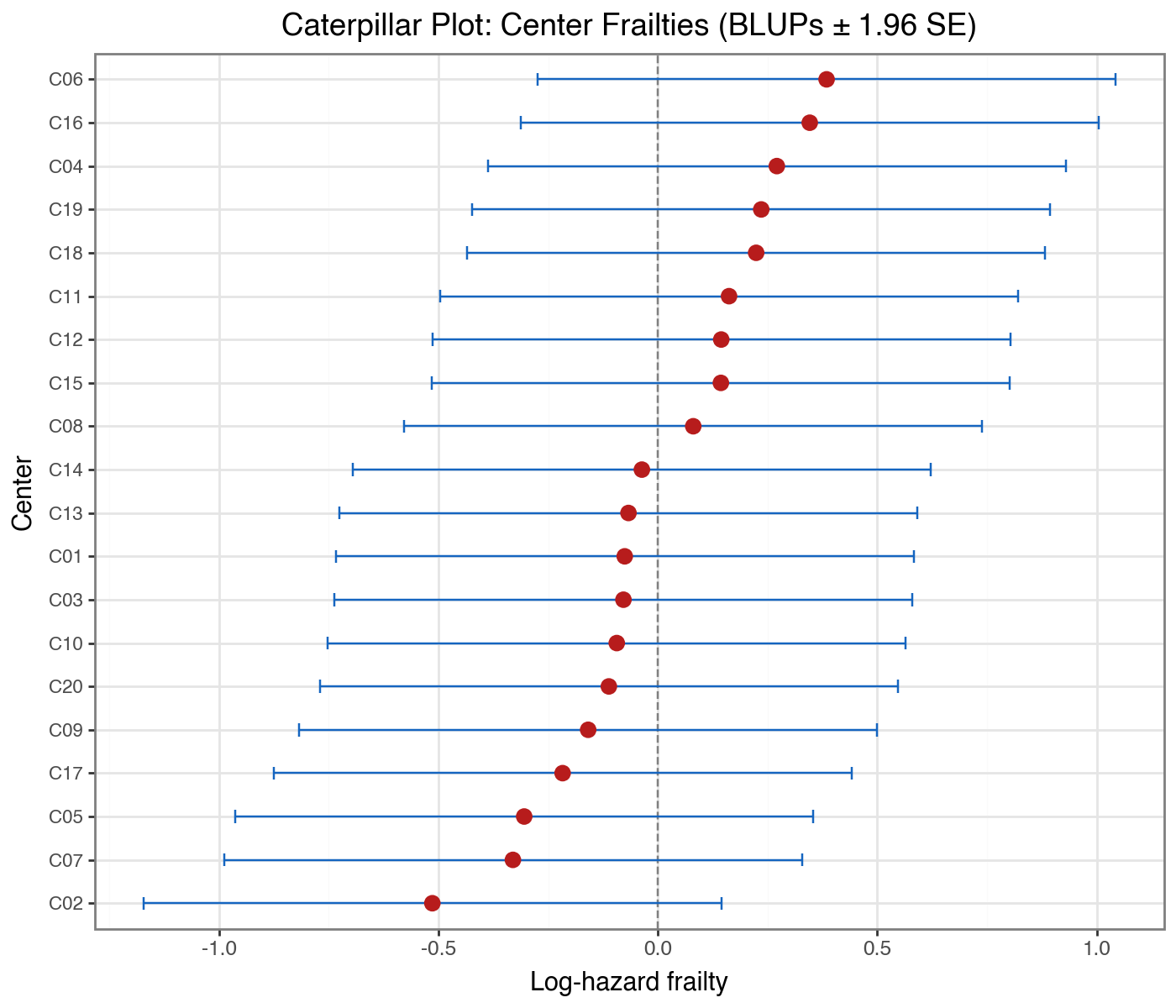
title='Caterpillar Plot: Center Frailties (BLUPs ± 1.96 SE)',
x='Center',
y='Log-hazard frailty',
)
+ theme_bw()
+ theme(figure_size=(7, 6), axis_text_y=element_text(size=8))
)
bh = result.baseline_hazard
(
ggplot(bh, aes(x='time', y='hazard'))
+ geom_step(color='#1565C0', size=0.8)
+ labs(
title='Breslow Cumulative Baseline Hazard',
x='Time (months)',
y='Cumulative baseline hazard H\u2080(t)',
)
+ theme_bw()
+ theme(figure_size=(8, 4))
)
---------------------------------------------------------------------------
NameError Traceback (most recent call last)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/mapping/evaluation.py:165, in evaluate(aesthetics, data, env)
164 try:
--> 165 new_val = env.eval(col, inner_namespace=data)
166 except Exception as e:
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/mapping/_env.py:71, in Environment.eval(self, expr, inner_namespace)
70 code = _compile_eval(expr)
---> 71 return eval(
72 code, {}, StackedLookup([inner_namespace] + self.namespaces)
73 )
File <string-expression>:1
NameError: name 'hazard' is not defined
The above exception was the direct cause of the following exception:
PlotnineError Traceback (most recent call last)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/IPython/core/formatters.py:1036, in MimeBundleFormatter.__call__(self, obj, include, exclude)
1033 method = get_real_method(obj, self.print_method)
1035 if method is not None:
-> 1036 return method(include=include, exclude=exclude)
1037 return None
1038 else:
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/ggplot.py:172, in ggplot._repr_mimebundle_(self, include, exclude)
169 self.theme = self.theme.to_retina()
171 buf = BytesIO()
--> 172 self.save(buf, "png" if format == "retina" else format, verbose=False)
173 figure_size_px = self.theme._figure_size_px
174 return get_mimebundle(buf.getvalue(), format, figure_size_px)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/ggplot.py:681, in ggplot.save(self, filename, format, path, width, height, units, dpi, limitsize, verbose, **kwargs)
632 def save(
633 self,
634 filename: Optional[str | Path | BytesIO] = None,
(...) 643 **kwargs: Any,
644 ):
645 """
646 Save a ggplot object as an image file
647
(...) 679 Additional arguments to pass to matplotlib `savefig()`.
680 """
--> 681 sv = self.save_helper(
682 filename=filename,
683 format=format,
684 path=path,
685 width=width,
686 height=height,
687 units=units,
688 dpi=dpi,
689 limitsize=limitsize,
690 verbose=verbose,
691 **kwargs,
692 )
694 with plot_context(self).rc_context:
695 sv.figure.savefig(**sv.kwargs)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/ggplot.py:629, in ggplot.save_helper(self, filename, format, path, width, height, units, dpi, limitsize, verbose, **kwargs)
626 if dpi is not None:
627 self.theme = self.theme + theme(dpi=dpi)
--> 629 figure = self.draw(show=False)
630 return mpl_save_view(figure, fig_kwargs)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/ggplot.py:306, in ggplot.draw(self, show)
304 with plot_context(self, show=show):
305 figure = self._setup()
--> 306 self._build()
308 # setup
309 self.axs = self.facet.setup(self)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/ggplot.py:379, in ggplot._build(self)
375 layout.setup(layers, self)
377 # Compute aesthetics to produce data with generalised
378 # variable names
--> 379 layers.compute_aesthetics(self)
381 # Transform data using all scales
382 layers.transform(scales)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/layer.py:485, in Layers.compute_aesthetics(self, plot)
483 def compute_aesthetics(self, plot: ggplot):
484 for l in self:
--> 485 l.compute_aesthetics(plot)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/layer.py:269, in layer.compute_aesthetics(self, plot)
262 def compute_aesthetics(self, plot: ggplot):
263 """
264 Return a dataframe where the columns match the aesthetic mappings
265
266 Transformations like 'factor(cyl)' and other
267 expression evaluation are made in here
268 """
--> 269 evaled = evaluate(self.mapping._starting, self.data, plot.environment)
270 evaled_aes = aes(**{str(col): col for col in evaled})
271 plot.scales.add_defaults(evaled, evaled_aes)
File ~/repos/gpg/interlace/.venv/lib/python3.14/site-packages/plotnine/mapping/evaluation.py:168, in evaluate(aesthetics, data, env)
166 except Exception as e:
167 msg = _TPL_EVAL_FAIL.format(ae, col, str(e))
--> 168 raise PlotnineError(msg) from e
170 try:
171 evaled[ae] = new_val
PlotnineError: "Could not evaluate the 'y' mapping: 'hazard' (original error: name 'hazard' is not defined)"
<plotnine.ggplot.ggplot object at 0x12668c750>
profiles = pd.DataFrame({
'treatment': [0, 0, 1, 1],
'age_c': [-1.0, 1.0, -1.0, 1.0], # ±10 years from mean (age 48 / 68)
'time': [1.0, 1.0, 1.0, 1.0],
'event': [0, 0, 0, 0],
'center': ['C_new', 'C_new', 'C_new', 'C_new'],
'label': ['Standard, age 48', 'Standard, age 68',
'New, age 48', 'New, age 68'],
})
profiles['predicted_HR'] = result.predict(
newdata=profiles,
type='risk',
include_re=False,
)
print("Predicted hazard ratios relative to baseline (age 58, standard treatment):")
print(
profiles[['label', 'predicted_HR']]
.rename(columns={'label': 'Profile', 'predicted_HR': 'HR (no frailty)'})
.to_string(index=False)
)
Predicted hazard ratios relative to baseline (age 58, standard treatment):
Profile HR (no frailty)
Standard, age 48 0.906359
Standard, age 68 1.103315
New, age 48 0.492626
New, age 68 0.599676
Summary¶
Results¶
Parameter |
True |
Estimated |
HR (est.) |
|---|---|---|---|
|
−0.431 |
(see table above) |
~0.65 |
|
0.223 |
(see table above) |
~1.25 |
Center SD |
0.500 |
(see table above) |
— |
Workflow checklist¶
[x] Generate synthetic clustered survival data with known parameters
[x] Exploratory KM curves and center heterogeneity
[x] Fit
interlace.coxme()with Gaussian frailty[x] Inspect hazard ratios and compare to ground truth
[x] Forest plot for coefficient summaries
[x] Caterpillar plot for center BLUPs
[x] Breslow baseline cumulative hazard
[x] Predictions for new patients (unseen center → frailty shrinks to 0)
See also¶
Cox frailty quickstart — minimal working example and API overview
coxme — full API reference